..and continuing in the thread of long-overdue R package updates, we’ve got a new ndtv version out as well.
Peek into the time prism
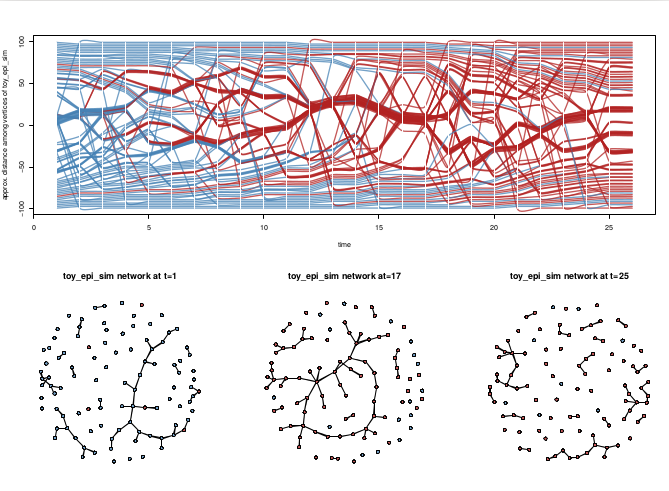
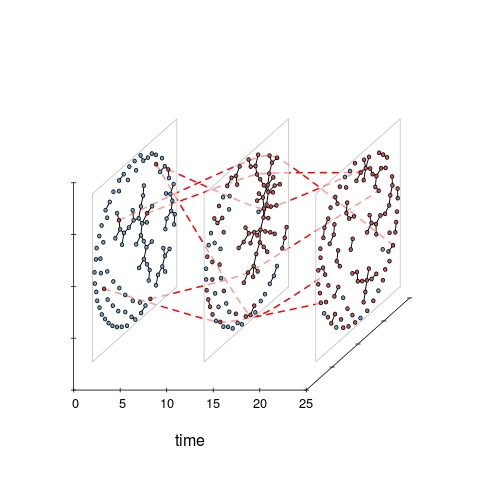
The package release adds some “whiteboard candy”: 2.5D orthogonal projection of networks in time along a z axis. For lack of a better name, I’ve dubbed it a timePrism (let me know if you find a pre- existing better name). Think of viewing all of the slices from a filmstrip from an angle. Probably hard to follow for large networks (or lots of time slices) but nice for illustrating concepts in temporal networks when you want to convey time and structure and can accept some loss of detail. Especially with the ability to include splines connecting specific vertices for highlighting trajectories.
library(ndtv)
data(toy_epi_sim)
timePrism(toy_epi_sim,
orientation=c('z','y','x'), # swap axes
angle=40,
spline.v=c(7, 29, 36, 70, 82, 96), # hilite the infected
spline.col='red',
spline.lwd=2,
box=FALSE,
planes=TRUE, # draw a semi-transparent 'plane' under each net
vertex.col='ndtvcol' # use pre-created infection color scheme)
Continue reading new ndtv release: 0.7 →